We neuroscientists tend to make a lot of assumptions about the brain: First, that everybody has one; second, that the more your neurons fire, the more blood will flow to those neurons; and third, that from the first two assumptions we can create sophisticated, impressive-sounding models and theories about the brain that explain everything, from consciousness and being to schizophrenia and Nutella addiction. Not that we aren't humble about it.
However, one of the most tenuous, sometimes ridiculous notions we have is that this blood flow, this continuous lifestream attempting to quench the brain's unslakable thirst with each pulse, is the same, acts the same, looks the same in every nook and cranny, every bump and crevice, every gyrus and sulcus of cortical and subcortical real estate. Why would we maintain such a clearly psychotic fantasy in the first place? Assumptions, madame, pure assumptions.
A word or two is in order about the origins of this one assumption in particular. In the beginning, a band of intrepid scientists observed that blood flow changes in primary sensory areas resulted in a gamma-shaped curve that peaks around five seconds followed by a long-lasting decline and undershoot around the thirty-second mark; at high field strengths, sometimes a small initial undershoot can also be observed. The results were so consistent in these areas, and the effects so strong, that these observations were soon cast into a model called the canonical hemodynamic response function (HRF), which has been widely used ever since.
And, for the most part, it has done pretty well. Typically a young, ambitious researcher, fueled by nothing more than the lust for knowledge and fame, for girls and gold and hazelnut spread - possibly even a swimming pool with all of the above mixed in there - will carry out his analyses using a whole-brain approach; that is, where the onset of each condition in each voxel is convolved with the hemodynamic response function. The height of the HRF is then estimated for each condition and averaged across trials, and contrasts can be carried out on these estimated heights. So far, so good; and plenty of high-quality research has been done using just this technique.
However, this is only a single parameter we are estimating for each condition; and if you happen to be a nerd, or a masochist, or a person who likes to do things the hard way - in other words, if you are in academia - you will doubtless be curious about other parameters you can harvest from your data. Sure enough, likeminded nerds have made these options available in the form of Finite Impulse Response (FIR) functions, which do not make any assumptions about the shape of the neural activity following a stimulus onset, and can therefore provide more detailed, flexible information about what is going on. All you do is specify that you wish to use that type of basis function, as well as the length of the interval you are interested in and how many time points you wish to estimate. Usually, FIR analyses are most interpretable when specified over a timecourse that doesn't include significant overlap with another condition, and when the timecourse is partitioned into units that are the same length as the time it takes to acquire each scan.
For now, I will focus on how to do this in AFNI. AFNI's version of FIR functions includes several ways to estimate each timepoint after a specified onset, including TENT, CSLPIN, and SIN basis functions, although for most purposes you will use TENT. (The TENT function uses a piecewise linear spline method to estimate brain activity at each timepoint; the details of this method are pretty complicated, which is a classy way of saying that I don't completely understand it.) The TENT function takes in three arguments: The beginning and ending times relative to the onset of a stimulus, or whatever timepoint you are interested in; and how many parameters to estimate. If b is the beginning and c is the end, with n being the number of parameters to estimate, then the time interval is (c-b)/(n-1). For example, if I wanted to estimate 5 parameters within a window from 0-8 seconds after a specified time, then the time interval between each point would be (8-0)/(5-1) = 2.
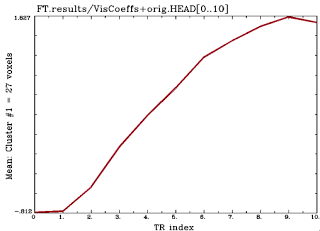
Although this would be usually implemented with event-related designs, I'll stick with AFNI's dataset #6, which uses a block design. Note that some of the shapes of the HRF can look pretty funky, compared to what you would expect when convolving with a canonical HRF; this should increase the already profound feelings of despair you have about cognitive neuroscience, and spur you to search for fields of more meaningful endeavor, such as specializing in pottery, martial arts, or poetry slams; specifically, slamming the country of Chile.
Here is what the code would look like for a typical FIR analysis in AFNI, assuming (har!) that you have already analyzed dataset #6:
#!/bin/tcsh
setenv subj = 'FT'
3dDeconvolve -input pb.$subj.r*.scale+orig.HEAD
-polort 3
-num_stimts 2
-stim_times 1 stimuli/AV1_vis.txt 'TENT(0,20,11)'
-stim_label 1 Vrel
-stim_times 2 stimuli/AV2_aud.txt 'TENT(0,20,11)'
-stim_label 2 Arel
-x1D X.xmat.1D -xjpeg X.jpg
-cbucket cstats.$subj
More information coming soon, but right now I have a bus to catch back to warm, sunny Minnesota! At least that's how my online travel agent, Amir, described it before I paid him five thousand dollars to arrange my trip and give me advice on how to woo the locals. Apparently, bright orange ski parkas are in style. Ladies?